Platform
Neomorph’s next-generation platform differs from the typical drug discovery workflow. Our access to hundreds of E3 ligases and multiple proprietary degrons is proof.
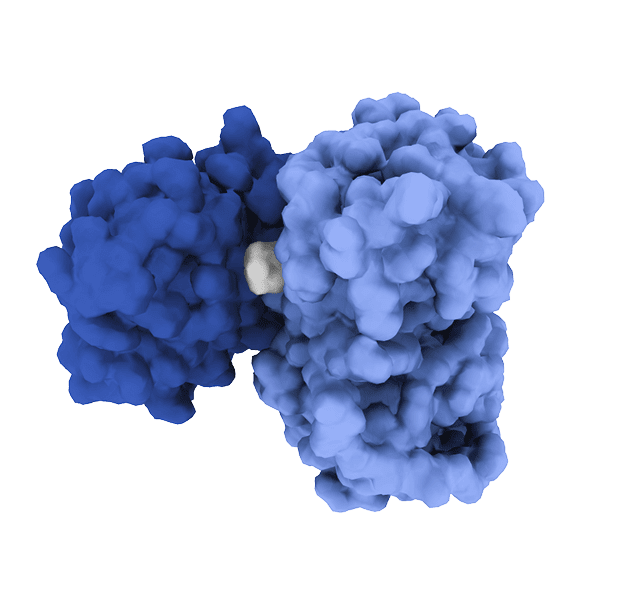
Molecular Glues
Some proteins have well-defined pockets. Others function through broad protein–protein interfaces, lack stable structure, or reside in intracellular compartments that are difficult for small molecules to access. For these reasons and more, roughly 85% of the human proteome remains out of reach using traditional small-molecule drugs and is considered “undruggable.”
Molecular glue degraders could reduce that percentage to zero.
These small-molecule drugs bypass the need for traditional binding pockets, offering a differentiated approach to drug discovery and the potential to target a broad swath of the proteome. With their low molecular weight, independence from ligand-binding sites, and broad proteome utility, molecular glue degraders are poised to break open the druggable genome. Approved drugs pomalidomide and indisulam validate the molecular glue mechanism, demonstrating that small molecules can repurpose E3 ligases for targeted protein degradation.
How Do Molecular Glues Work?
1. Bind an E3 ligase
Molecular glue drugs attach to an E3 ubiquitin ligase, a part of the cell’s protein-degradation machinery.
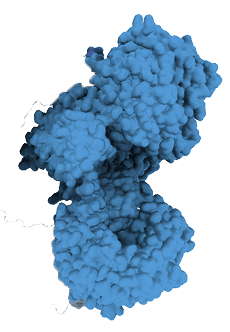
2. Create a new (“neomorphic”) binding surface
This binding reshapes the ligase, forming a new or “neomorphic” surface that can engage proteins that the E3 ligase may not normally recognize.
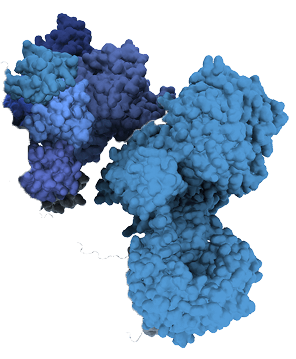
3. Recruit and eliminate disease-causing proteins
The neomorphic E3 ligase recruits the target protein to the ligase, where it is tagged with ubiquitin for degradation and subsequently destroyed by the proteasome. The structural feature on the target protein that binds the ligase is called the degron.
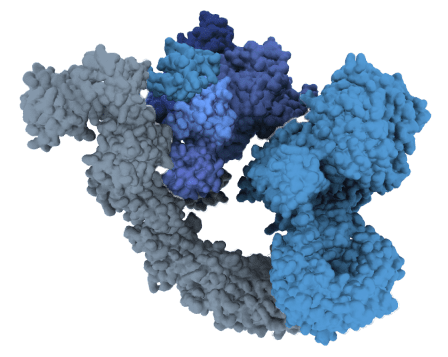
Beyond Cereblon
Identifying and designing new molecular glues has been historically challenging. This is why much of the field is singularly centered around cereblon: an E3 ligase capable of targeting roughly 3000 proteins. While impactful, that leaves more than 10,000 proteins unaddressed, and more importantly, countless patients waiting for new therapies.
To pursue these remaining targets, the field requires access to more than a single ligase. Neomorph is meeting that need.
Members of our leadership team played a central role in early cereblon discoveries, including solving the structure of the ligase bound to thalidomide and defining the first neosubstrate degron. We are applying our deep, foundational expertise to break new ground—manipulating cereblon in novel ways while exploring the more than 600 ligases that exist.
Today, we have created the largest known proprietary target space for molecular glues, including numerous proprietary degrons and an expansive number of enabled E3 ubiquitin ligases. We accomplish this with our next-generation molecular glue discovery platform.

The Glue That Makes It Work
The Neomorph platform ensures molecular glues are discovered via rational design rather than random chance. By combining multiple factors—ligase, degron, and chemical diversity; structural insights; and artificial intelligence—we pursue rational prospective drug discovery in a target-centric manner.
Custom-built tools allow us to design glue libraries, synthesize candidates, test activity, identify novel neosubstrates and ultimately validate them. This approach has generated a rich pipeline of potential molecular glue degraders, positioning us to rapidly advance candidates across multiple disease areas.

Ligase Diversity
- Expansive set of enabled novel ligases
- Abundant proprietary molecular glue systems
Ligase Diversity
- Expansive set of enabled novel ligases
- Abundant proprietary molecular glue systems
We are pursuing both actionable near-term opportunities in proven systems, such as in the canonical cereblon degron and our proprietary alternative degrons, in addition to next-generation mechanisms to uncover hundreds of additional ligases with potential for glue activity for massive expansion of the druggable genome.
The Neomorph ligase database integrates biological, structural and chemical data with ranking based on proprietary criteria. These include AI methods to identify “gluable” interfaces, help select ligases, and assess compatibility of ligases for potential neo-substrates, as well as biophysical parameters to design small molecules directed to enabling protein-protein interactions.

Degron Diversity
- Multiple novel degrons discovered
- Expanding druggable genome
Degron Diversity
- Multiple novel degrons discovered
- Expanding druggable genome
We have identified multiple novel degrons beyond the canonical G-loop motif—expanding proteome coverage. Our structure-based AI predicts portability across a significant portion of the human proteome.

Chemical Diversity
- Proprietary glue library optimized for neomorphic activity
- Broad proteome coverage
Chemical Diversity
- Proprietary glue library optimized for neomorphic activity
- Broad proteome coverage
The Neomorph glue library is optimized for broad degradative coverage of the proteome and chemical space while retaining excellent drug-like properties. We have additional screening capabilities using our proprietary DEL libraries, which allow for the direct discovery of novel glue degraders. Our approach enables screening millions of molecular glue candidates to identify diverse and new chemotypes, resulting in hits to high-value undruggable targets.

Structural Insights
- Programs fully structurally enabled
- Rapid determination of complex structures
Structural Insights
- Programs fully structurally enabled
- Rapid determination of complex structures
We maintain world-class structural biology for novel glue discovery—a critical differentiator for rational drug design and the advancement of novel glue systems. Combining ubiquitin ligase protein chemistry, X-ray crystallography, and a world-class cryo-EM team, we have ternary complex structures for all Neomorph pipeline programs from the earliest stages of a project.

AI Capability
- Interface compatibility prediction
- Predictive glue design
AI Capability
- Interface compatibility prediction
- Predictive glue design
Our cutting-edge data science supports our innovation in the degradation space. We develop proprietary algorithms and integrate bioinformatics, computational chemistry, and machine learning across all platforms and programs.
Foundational Publications
PARTNER WITH US
.We’re constantly exploring partnerships that advance our work across the proteome. To collaborate or learn more about our technology, get in touch today.